Sophisticated free app for molecular visualization
"A must for any scientist working with porous materials"
This month the unique advanced iRASPA app has been launched. This app visualizes molecular structures of various types of porous materials with which material research can be facilitated and improved. In addition, it can help teachers with their explanation to students. The development of the iRASPA is done by a team of computational chemists: Dr David Dubbeldam (University of Amsterdam, Van 't Hoff Institute for Molecular Sciences), Sofia Calero (Universidad Pablo de Olavide in Seville, Spain) and Thijs Vlugt, Professor of Engineering Thermodynamics at the Process & Energy (3mE). Thijs Vlugt has contributed to the software model that uses algorithms to show and move molecular 3D structures. The coming period Vlugt will focus on the application to convert the images in iRASPA into high-quality film.
iRASPA
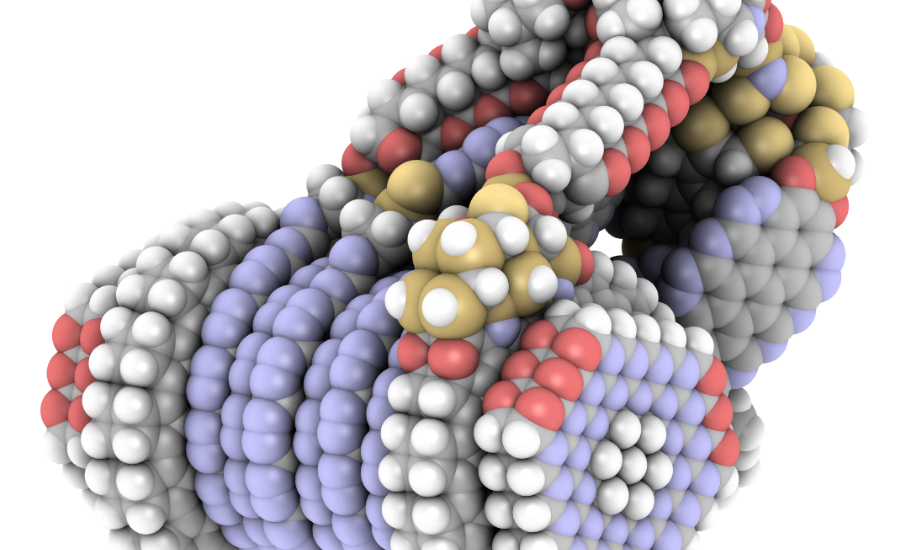
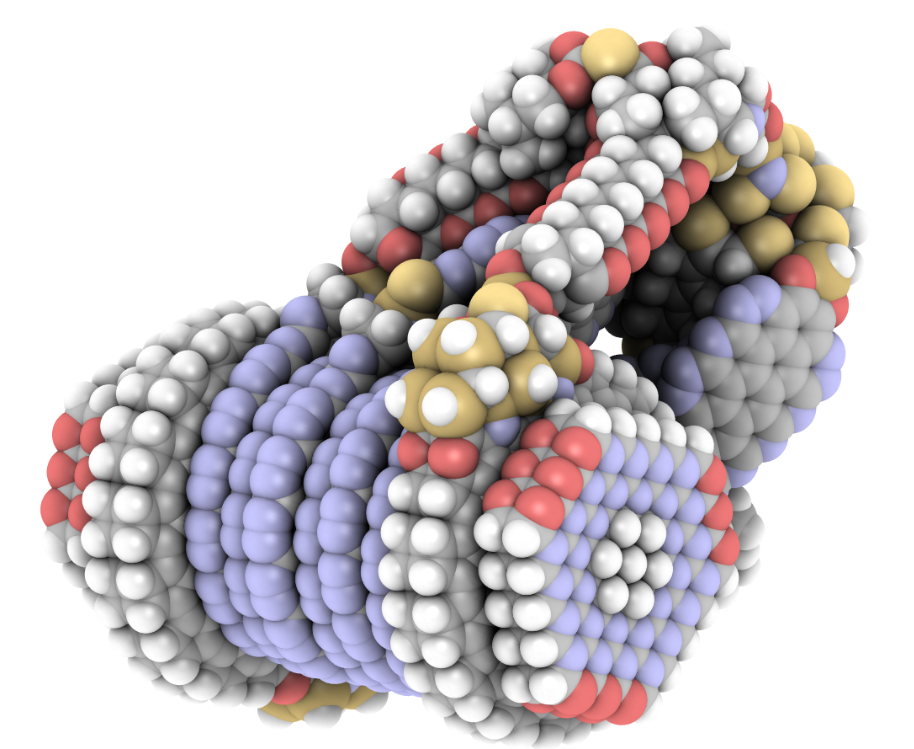
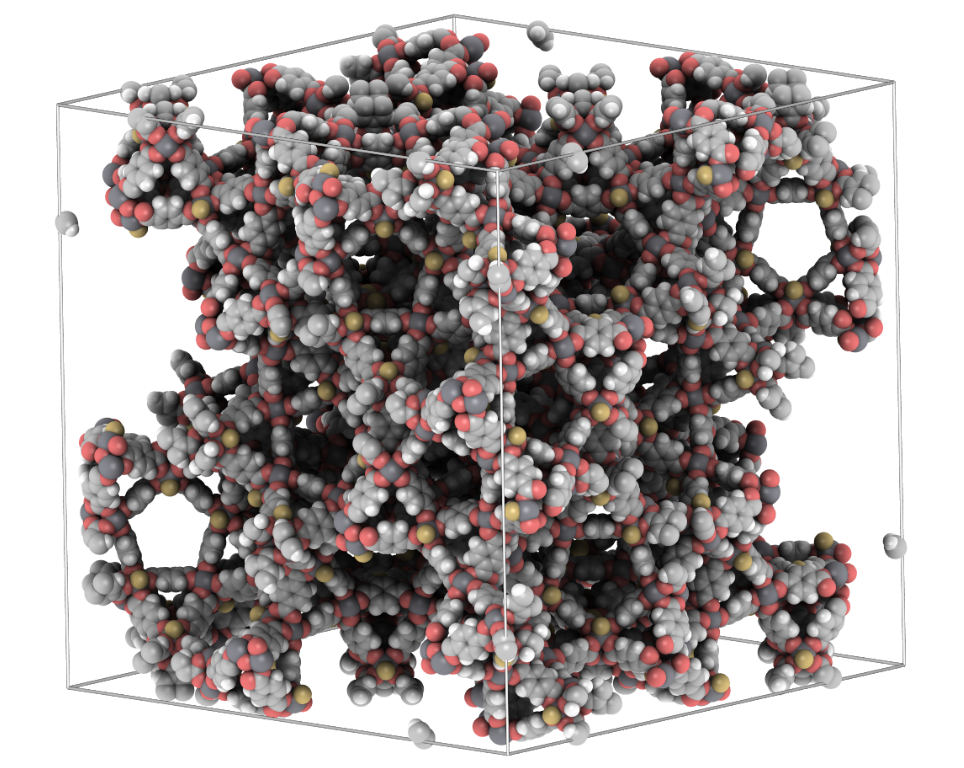
iRASPA is a molecular visualization package with editing capabilities aimed at materials science. Controlled by means of a versatile user interface it produces high quality images of complex molecules, metals, metal-oxides, ceramics, biomaterials, zeolites, clays, and metal-organic frameworks.
iRASPA is exclusively for macOS and as such can leverage the latest visualization technologies with stunning performance. Do you work with porous materials and are you curious what iRASPA can mean for your communication? Sign up now for the workshop!
iRASPA extensively utilizes GPU computing. For example, void-fractions and surface areas can be computed in a fraction of a second for small/medium structures and in a few seconds for very large unit cells. It can handle large structures (hundreds of thousands of atoms). It is much easier to see depth in the molecular structure by using an ambient occlusion scheme.
Acces to database
Via iCloud integration, iRASPA has access to the CoRE Metal-Organic Frameworks database containing approximately 8000 structures. All the structures can be screened (in real-time) using user-defined predicates. Structures can be selected for surface areas, void fraction, and other pore structure properties.
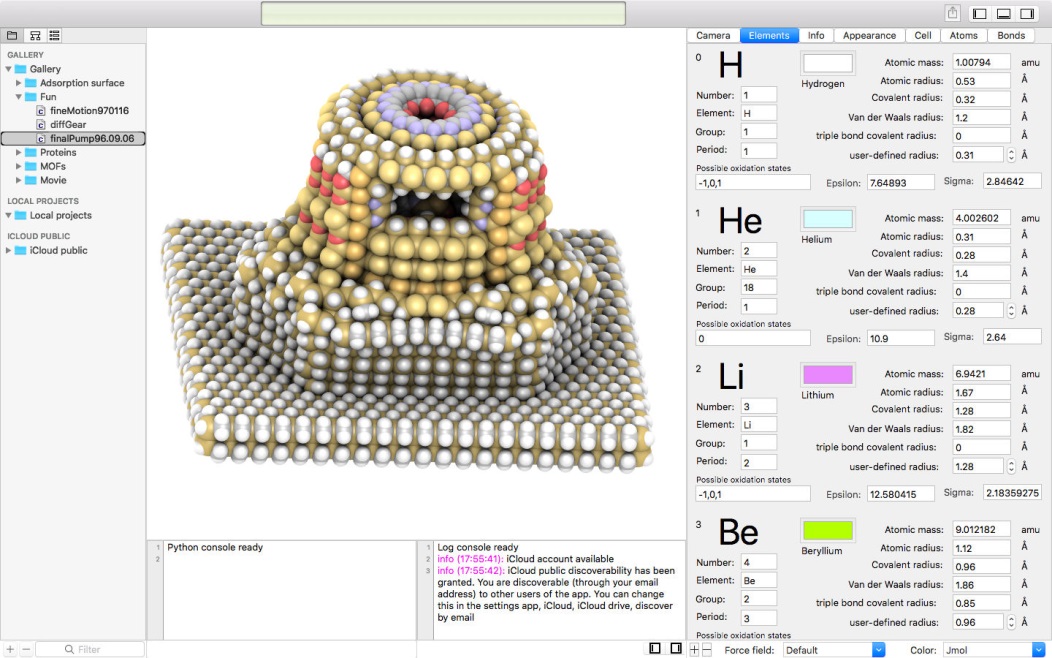
Main features of iRASPA are:
• structure creation and editing;
• creating high-quality pictures and movies;
• ambient occlusion and high-dynamic range rendering;
• collage of structures;
• (transparent) adsorption surfaces;
• text-annotation;
• cell replicas and supercells;
• symmetry operations like space group and primitive cell detection;
• screening of structures using user-defined predicates;
• GPU-computation of void-fraction and surface areas in a matter of seconds.
iRASPA has been created by David Dubbeldam (University of Amsterdam, The Netherlands), Sofia Calero (Universidad Pablo de Olavide, Seville, Spain) and Thijs Vlugt (Delft University of Technology, The Netherlands) with contributions from Randall Q. Snurr (Northwestern University, Evanston, USA).
A detailed description of iRASPA and its use is presented in a recent open-access article in Molecular Simulation: David Dubbeldam, Sofía Calero & Thijs J.H. Vlugt (2018) iRASPA: Molecular Simulation,
44: 8, 653-676, DOI: 10.1080 / 08927022.2018.1426855
- More visualization examples in the iRASPA Gallery.
- iRASPA website
- iRASPA at the App Store
- HIMS research group Computational Chemistry