Aster positioning
Yash Jawale
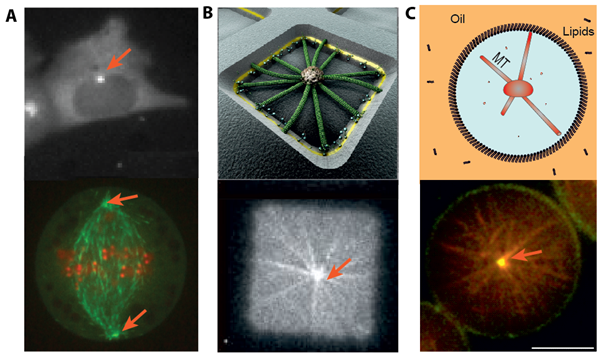
Important functions of eukaryotic cells, like motility or division, depend sensitively on cytoskeletal mechanics and organization. In particular, microtubules (MTs) are dynamic polymers that can move and position organelles such as their Microtubule-Organizing Center (MTOC), by pushing, pulling or sliding. For example, during migration CHO cells position their MTOC at the leading edge (Figure 1A, top), while during cell division the two MTOCs position at opposite poles, forming a spindle structure (Figure 1A, bottom).
Microtubules grow radially from their MTOCs. We aim to understand how one or two such radial arrays of microtubules can position in confinement and how this positioning depends on microtubule dynamics, confinement shape, forces and proteins known to be involved in positioning processes. For this, we adopt a bottom up approach by building in vitro minimal systems. We grow microtubules from nucleating centers and confine them in different geometries (Figure1 B and C), mimicking the confined environment of a cell. We add proteins known to affect the positioning of these MTOCs in vivo, either by influencing the dynamics of the microtubules or the forces exerted on the MTOCs.
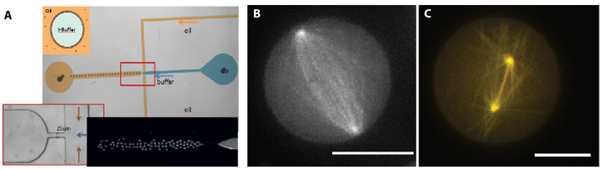
In our project ‘mitotic spindle’, we confine two dynamic asters inside 3D water-in-oil emulsion droplets made with microfluidics (Figure 2). We have chosen these 3D water-in-oil emulsion droplets for their spherical shape, which resembles a cell undergoing division. We study the positioning of these centrosomes by spinning disk confocal microscopy (Figure 2B and C + movie). We show that very simple systems, involving only microtubule dynamics, confinement, pushing and pulling forces can lead to self-organized patterns that are biologically relevant. In particular, we reproduce a ‘mitotic spindle’ like organization with just these components.
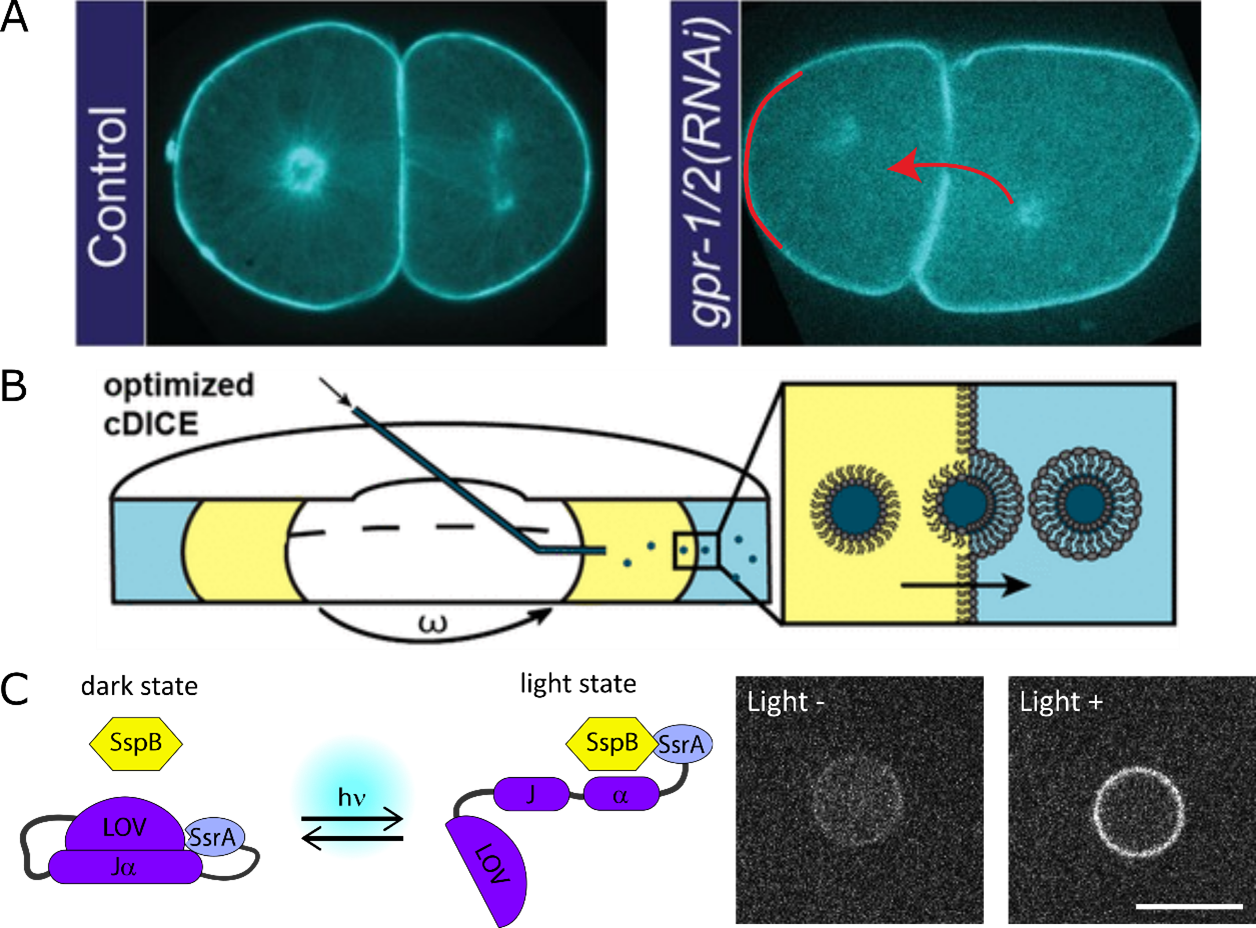
Cells also undergo asymmetric cell division for events like C. elegans developmental axis formation (Figure 3A left), stem cell homeostasis. Asymmetry is controlled by spindle positioning and can be altered by changing force generators’ (dynein) balance (Figure 3A right). We are now transferring the minimal spindle system into liposomes made using cDICE (Figure 3B). The force generators’ recruitment is controlled using photo-switchable proteins like iLID/SspB pair (Figure 3C).