Combination of microscopy techniques makes images twice as sharp
Researchers at Delft University of Technology have combined two existing super-resolution microscopy techniques to create a new method. Many experts thought that combining these techniques was not technically possible. The new, combined method enables researchers to visualize the tiny components of living cells better than ever before. Among other things, this can lead to new insights for healthcare.
Super-resolution microscopy is a groundbreaking technology that allows researchers to look inside living cells. The technique makes use of the fluorescent proteins that can be found in jellyfish and other creatures. Researchers can use gene editing to attach these fluorescent proteins to molecules.
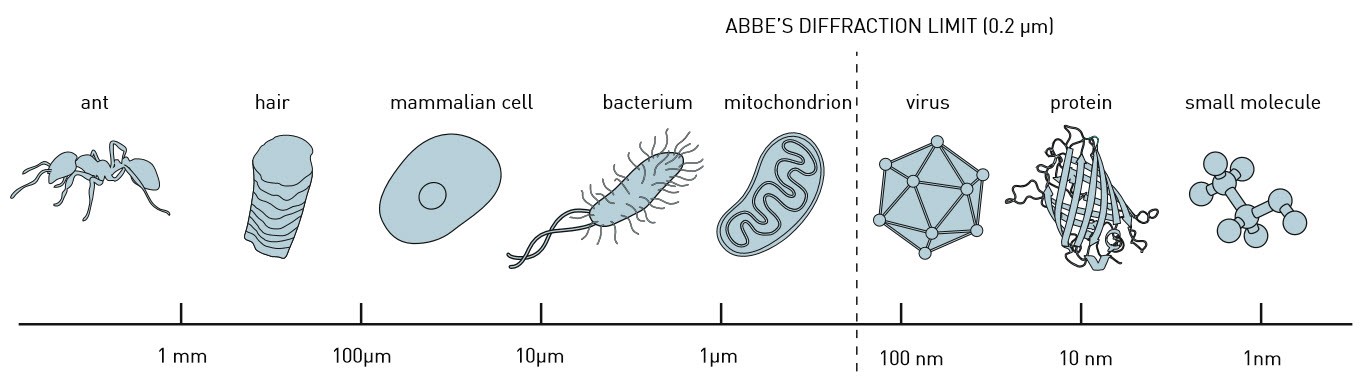
Shining a laser on such a protein causes it to emit a little bit of light a short while later. Sensitive sensors capture the light signals and, aided by algorithms that filter the valuable data out of the noise, researchers can use this data to build up a picture. The development of super-resolution microscopy has been an enormous leap forward. "With an ordinary optical microscope you can create images on a scale of about half a micron", says researcher Sjoerd Stallinga. "With super-resolution microscopy you can do that ten times better."
Further refining
The field of super-resolution microscopy has developed rapidly over the past decade. In 2008, three top researchers in the field were awarded the Nobel Prize in Chemistry for the discovery and further development of the aforementioned luminous protein, called GFP (Green Fluorescent Protein). In 2014, three other researchers received the Nobel Prize for the use of these proteins in what became known as 'superresolution microscopy'. Since then, experts from all over the world have been looking for ways to further refine the technique. It is a true race to the bottom.
There are currently two methods within super-resolution microscopy. In the first method, called Single Molecule Localization Microscopy (SMLM), researchers ensure that molecules are randomly switched on or off. "Most of the time they are off, but sometimes they light up," explains Stallinga. "When you make a video, you will see a glittering starry sky, where each star is a speck of light of one molecule. Since the different molecules are only switched on occasionally, you can determine their location very precisely. And if you analyse the entire film, you can make a reconstruction of the cell structure you are looking at."
Light modelling
SMLM's limitation is that only a few hundred up to a few thousand photons are released from the molecules. By way of comparison: the number of photons that hit your retina when you look at your own hand, is about 1,000,000,000,000,000,000. "The question we have asked ourselves is: how can we make even better use of the tiny amount of light that comes from the fluorescent molecules?", says researcher Carlas Smith.
That's where the second method in super-resolution microscopy comes in. With this method, Structured Illumination Microscopy (SIM) researchers model the laser light into a very finely striped pattern of light-dark-light-dark. They then project that pattern onto their sample. "If a molecule is in a bar of light, we excite it and it starts emitting photons, causing us to see it. If, however, the molecule is in a dark area, we don't see it," Smith explains. "By placing both a light pattern with horizontal and vertical stripes on top of the sample and moving the pattern back and forth, we can determine the location of molecules very precisely.
Three dimensions
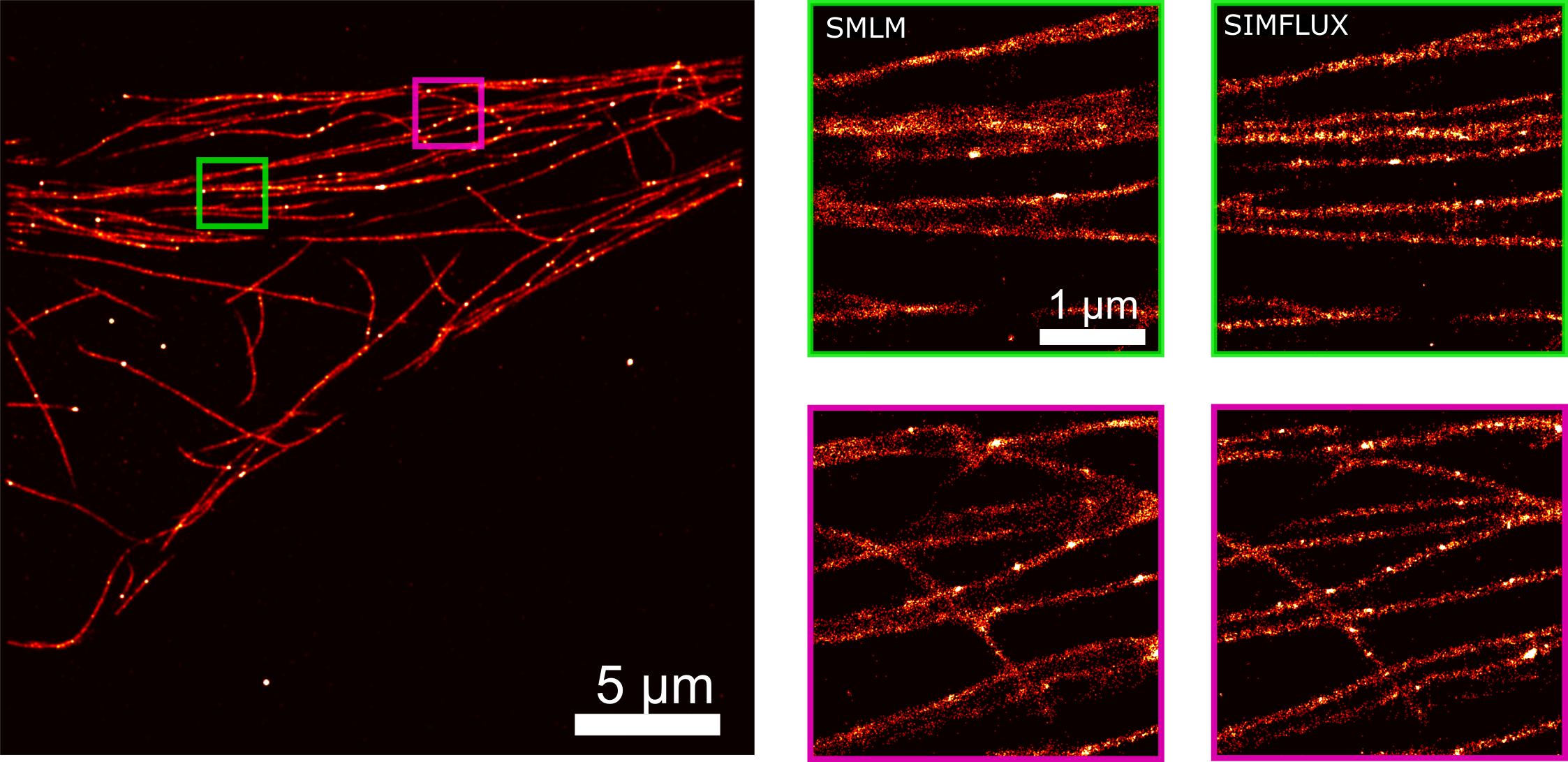
After a lot of theoretical and practical work, the Delft researchers succeeded in combining the two techniques, SMLM and SIM, in one set-up. They call their new technique SIMFLUX. In order to demonstrate that the method works, the researchers imaged an artificially produced DNA structure. "We spent another year to actually look inside a cell," says researcher Bernd Rieger. These efforts yielded a clear picture of the protein wires and tubes that together form the skeleton of a cell: the so-called cytoskeleton.
The new method is a major step forward. It allows researchers to zoom in on structures of between five and ten nanometres, about twice as good as was possible with the existing SMLM method. Is there still room for improvement? Probably so. "The logical next step would be to make images in three dimensions," says Rieger. "That's another big challenge, but we already have some ideas."
Interfaculty
This research was carried out at two faculties (TNW/Imphys and 3mE/DCSC) from start to finish. While the experiments at TNW were done in the Rieger/Stallinga group, the analysis software at 3mE was made in the Smith group. "The PhD candidates and postdocs working on the research from both groups crossed the Mekelweg at least 100 times for consultation," says Stallinga. "We have noticed that the Mekelpark is not an insurmountable obstacle to a successful collaboration.
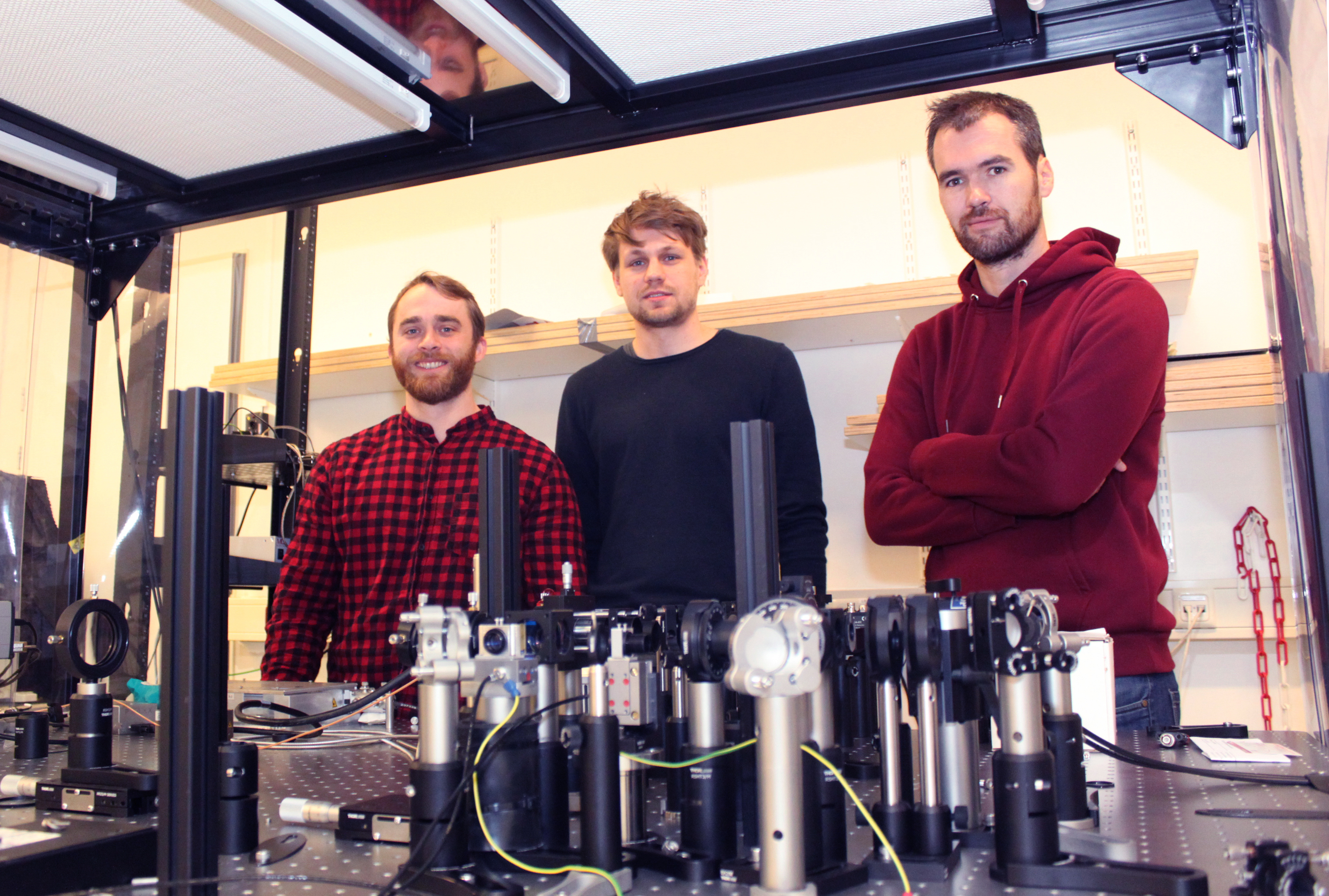
Postdoc Taylor Hinsdale (left), PhD candidate Rasmus Thorsen (middle) and PhD candidate Jelmer Cnossen (right) at the new microscopy setup.