Researchers shed new light on DNA replication
In preparation for cell division, cells need to copy (‘replicate’) the DNA that they contain. A team of researchers from TU Delft, collaborating with investigators from the Francis Crick Institute in London, has now shown that the protein building blocks involved in the initial steps of DNA replication are mobile but reduce their speed at specific DNA sequences on the genome. Their findings, which were published in the open-access journal Nature Communications on 26 March, were facilitated using an integrated approach involving biophysics and biochemistry that will propel new discoveries in the field.
DNA replication is fundamental for life – indeed, we copy a light-year’s worth of DNA over the course of our lifetimes. This copying must be carried out accurately by a molecular machine consisting of protein components, and as such it is carefully controlled. However, how this control of the protein components is achieved in the watery environment of the cell is by no means always clear.
Bottom-up approach
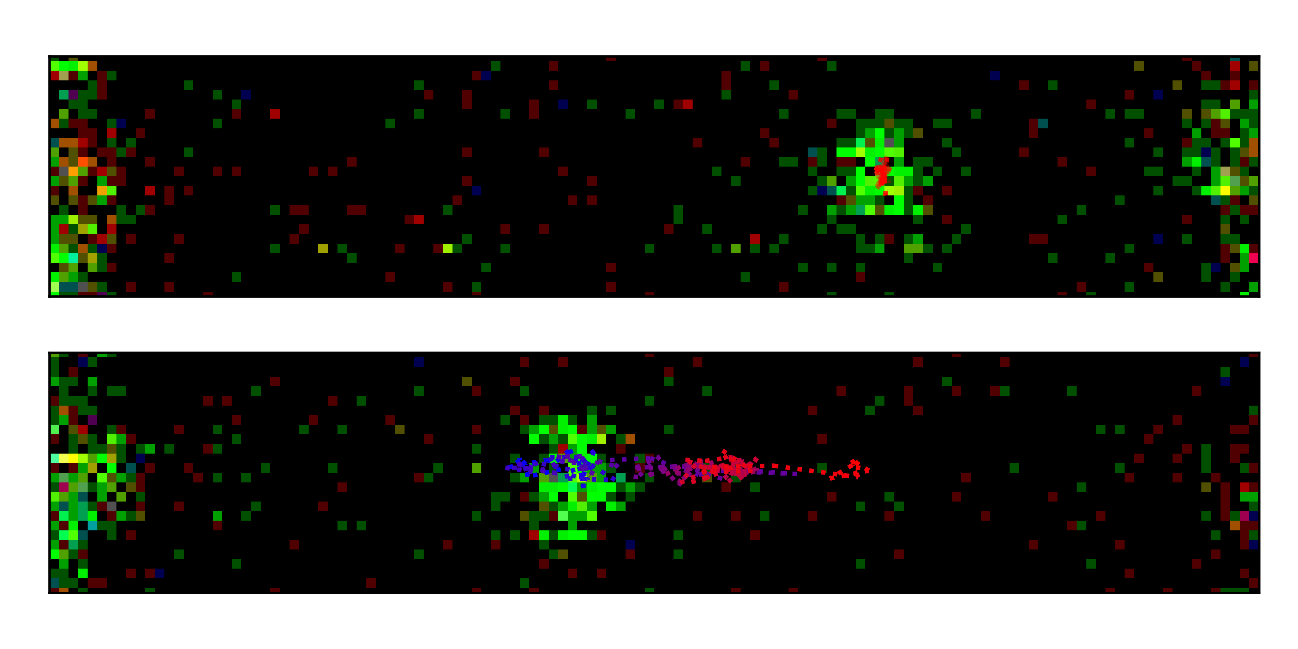
To study the dynamics involved in the first phase of DNA replication, the TU Delft team, led by Spinoza laureate Prof. Dr. Nynke Dekker, collaborated with Francis Crick Institute group leader Dr. John Diffley to implement a bottom-up approach in which the first part of the replication machinery was built up from its individual components on a single molecule of DNA held in place using optical tweezers. By attaching fluorescent labels to the proteins of interest, the researchers could then reveal their movement on the DNA.
In this way, the researchers established for the first time that the protein that sets into motion the building of the replication machinery (the origin recognition complex (ORC) – originally discovered by the winner of the 2020 Dr. H.P. Heineken Prize for Biophysics and Biochemistry, Dr. Bruce Stillman) is a mobile protein that diffuses along the DNA. Importantly, however, its motion is brought to a halt at particular DNA sequences that ORC was previously established to have high affinity for. This reduction of ORC mobility at these sequences then facilitates the assembly of additional components of the replication machinery at these locations on the DNA.
New light
Using this approach, the researchers also found that the key motor component of the replication machinery (known as MCM) can stably associate with the DNA not only in its canonical double hexameric form, but also in a hitherto unknown single hexameric form uncoupled from ORC that displays significant mobility. This expansion of knowledge about the forms that MCM adopts on DNA may imply that its role inside cells is larger than what textbook knowledge currently indicates.
With this work, the TU Delft and Francis Crick Institute researchers shed new light on a fundamental biological process that affects our daily lives through its connections to embryo development, tissue regeneration, and cancer progression. Quoting the first author of the work, Dr. Humberto Sánchez, “We are just at the beginning of a molecular journey that will be full of surprises.”

Prof. dr. N.H. Dekker
